The ExD AQ-25x Option
Practical Electron Fragmentation for Agilent
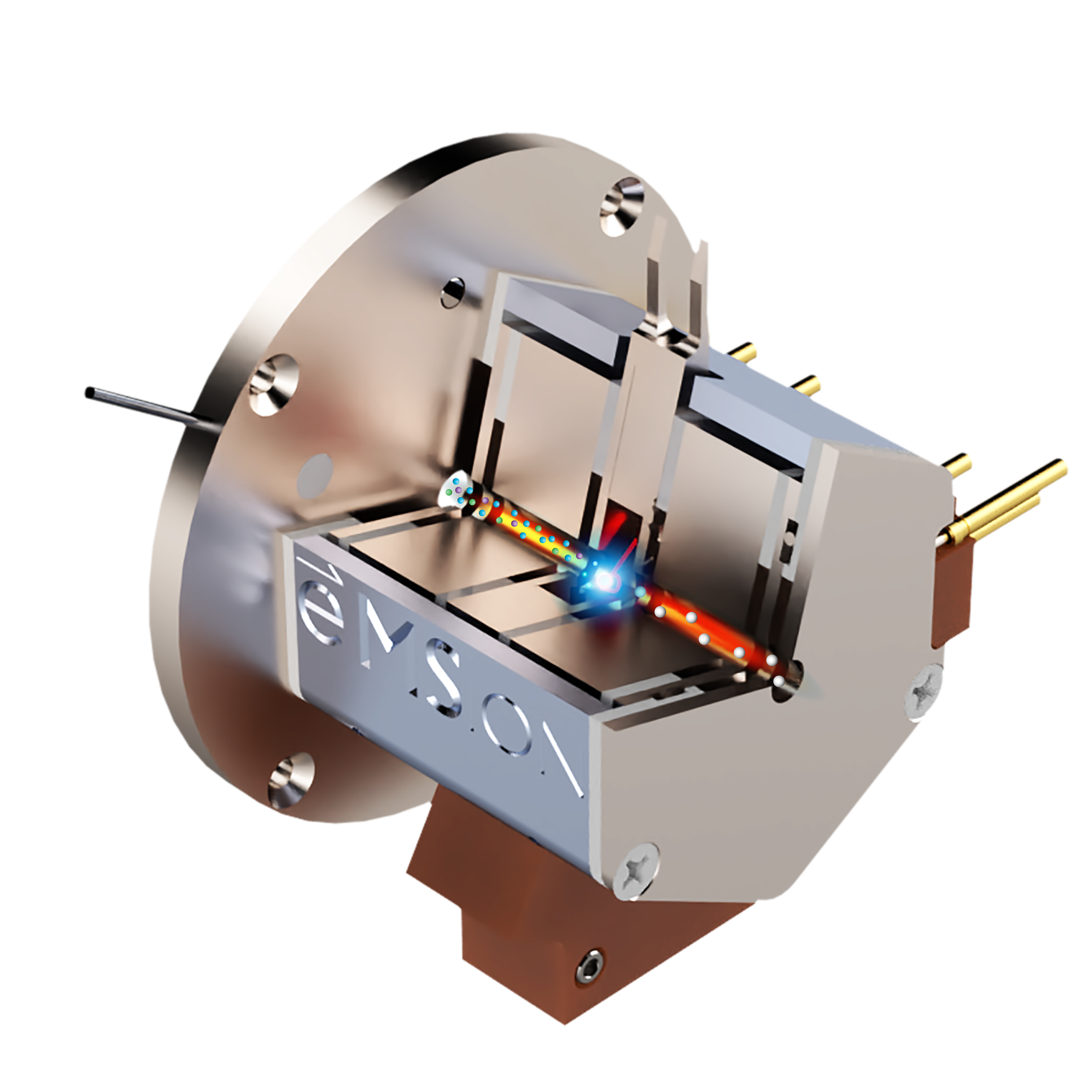

PERMANENT RING MAGNET
FILAMENT
ELECTROSTATIC LENSES
Add electron-based fragmentation with the ExD AQ-25x Option
Upgrades for Agilent high-resolution 6545XT AdvanceBio and 6560 Ion Mobility LC/Q-TOF are supported by Agilent teams in North America and Europe.
Applications
Increase sequence coverage of larger peptides and intact proteins beyond the limits of CID alone.
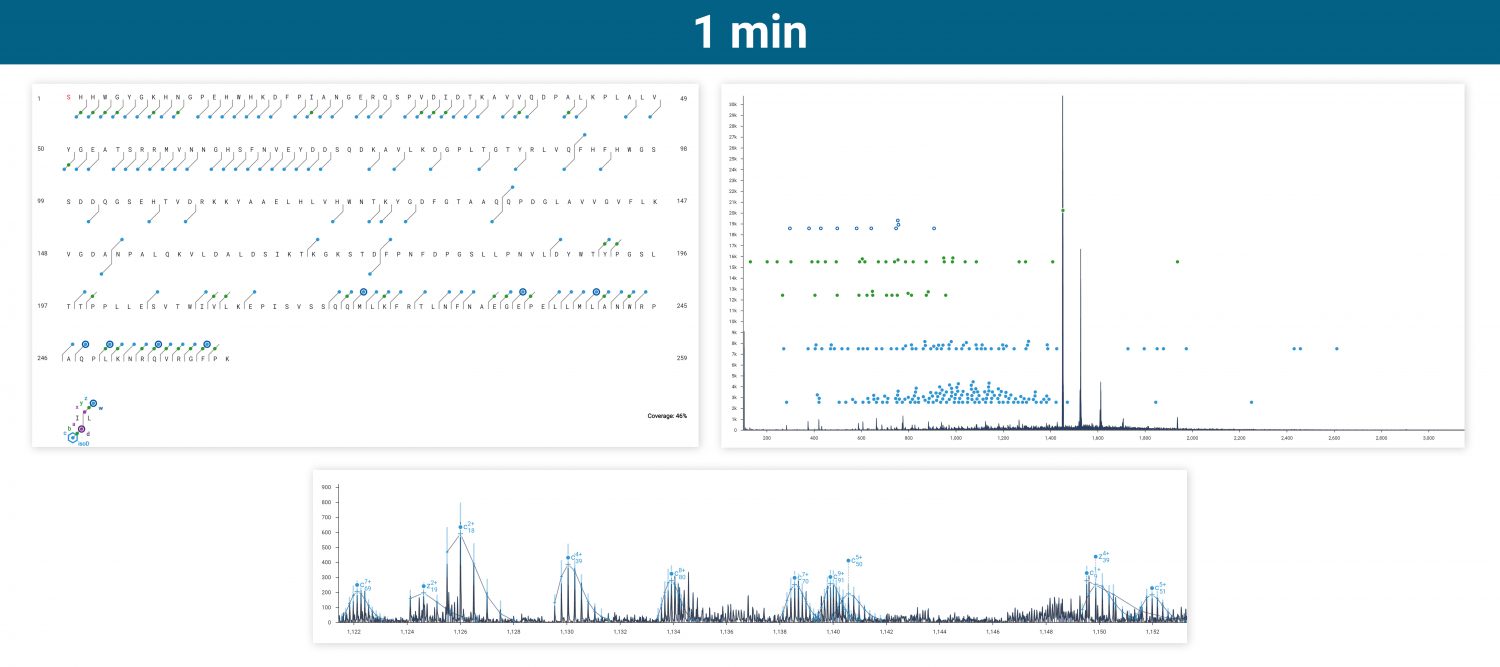
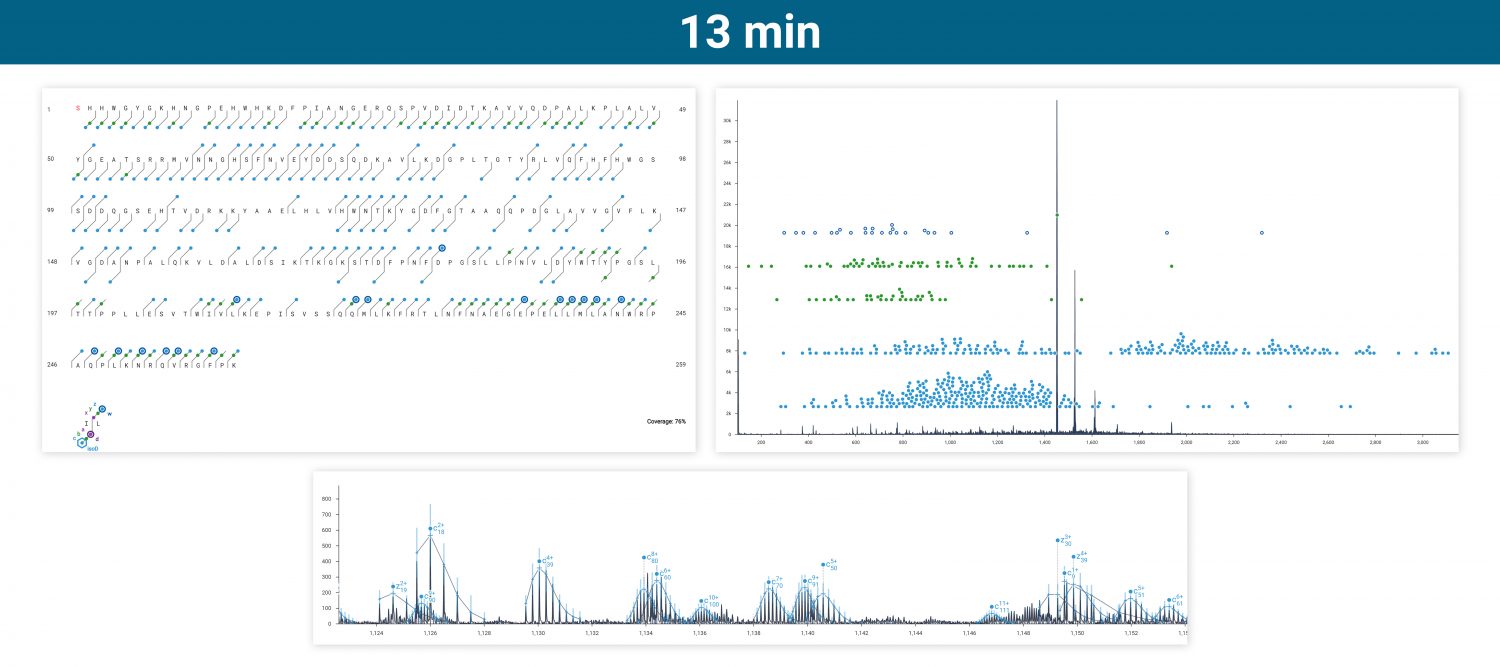
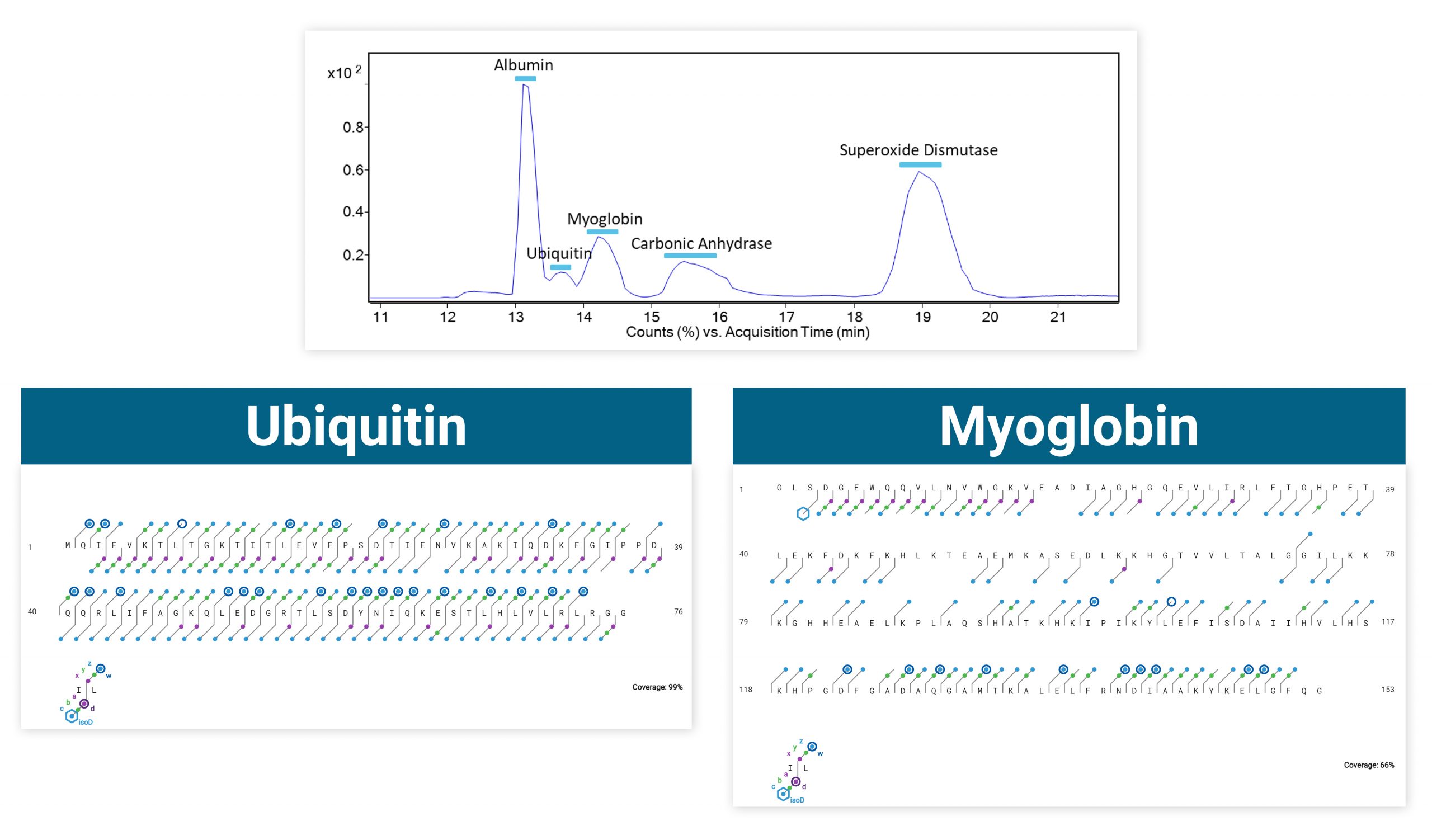
Intact carbonic anhydrase II was run via infusion on an Agilent 6545XT equipped with the ExD option. Top-down ECD was performed on the isolated 20+ charge state, producing amino acid sequence coverage of 28-76% depending on how many acquisitions were averaged.
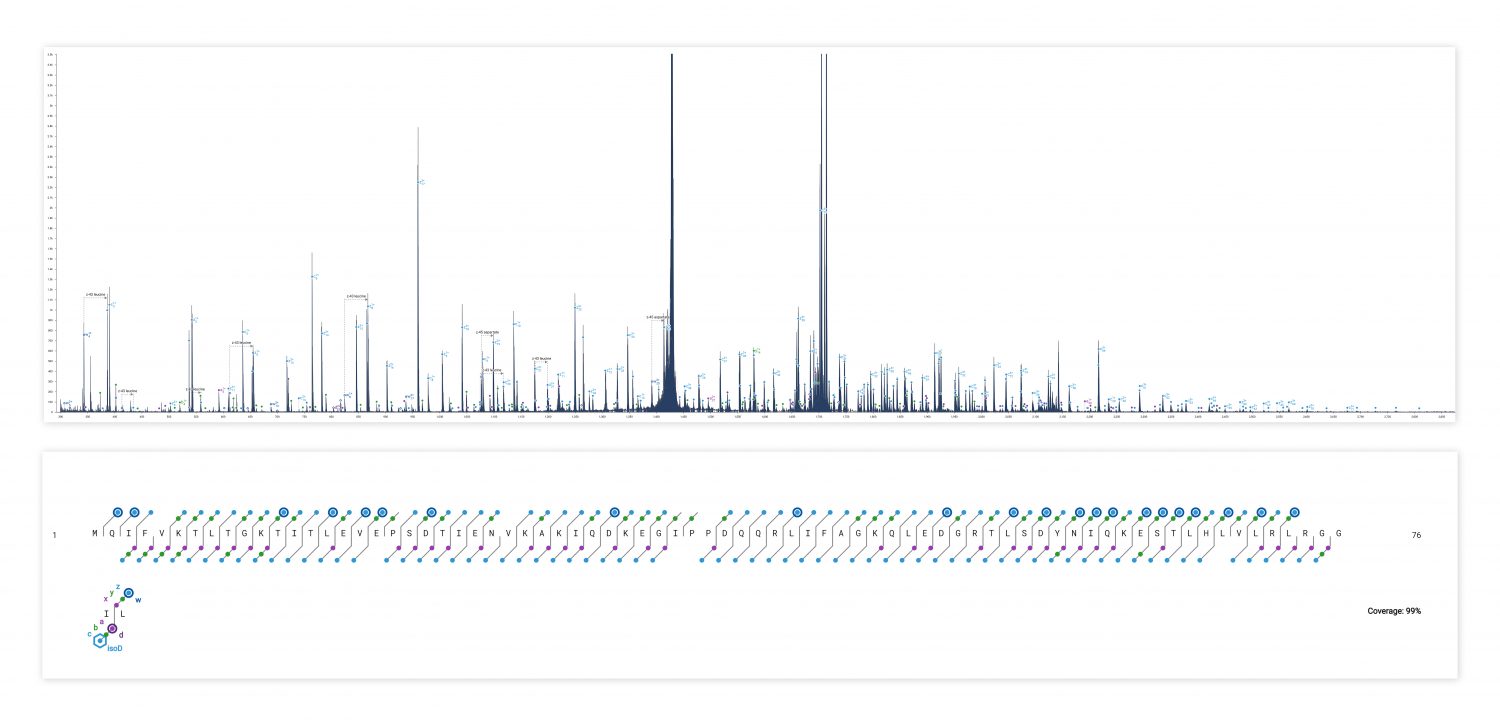
ECD produces sequence-informative fragments for proteins and large peptides while avoiding the non-informative fragmentation that can occur via CID or UVPD. Top-down ECD of 6+ ubiquitin in the 6545XT AdvanceBio LC/Q-TOF yielded a complete sequence of fragment ions, minus the N-terminus of proline residues, which are not dissociated by ECD.
Side chain fragments (w-ions) produced by ECD can be used to distinguish isobaric amino acids.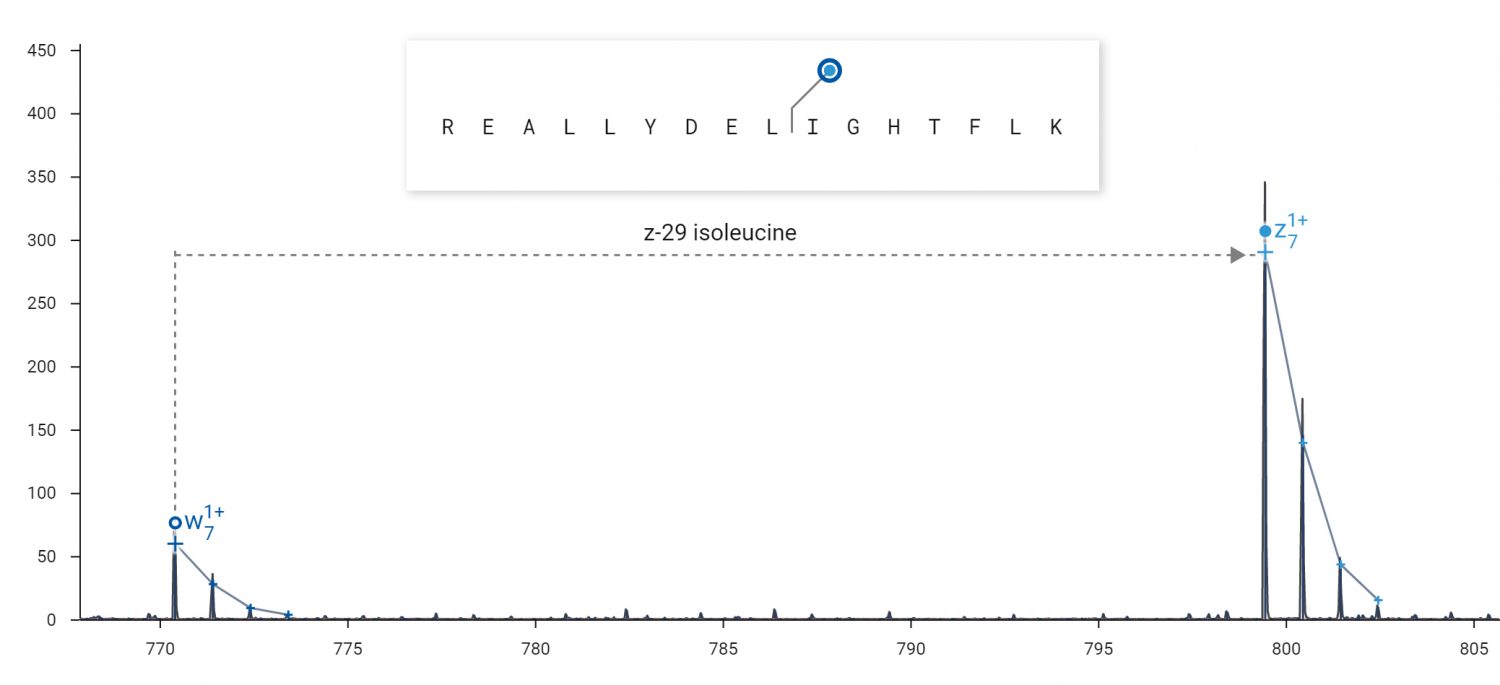

ECD can produce side-chain fragmentation useful for distinguishing leucine from isoleucine residues. ECD was performed on the 2+ peptide REALLYDELIGHTFLK in an Agilent 6545XT Q-TOF. Top: The diagnostic loss of 29 Da from the z7 fragment indicates the presence of isoleucine at residue 10. Bottom: The diagnostic loss of 43 Da from the z8 fragment indicates the presence of leucine at residue 9.
ECD fragmentation products unique to isoaspartate clearly differentiate it from the non-isomerized form. Isoaspartate is implicated in age-related protein inactivation and aggregation, and reduced efficacy of protein therapeutics.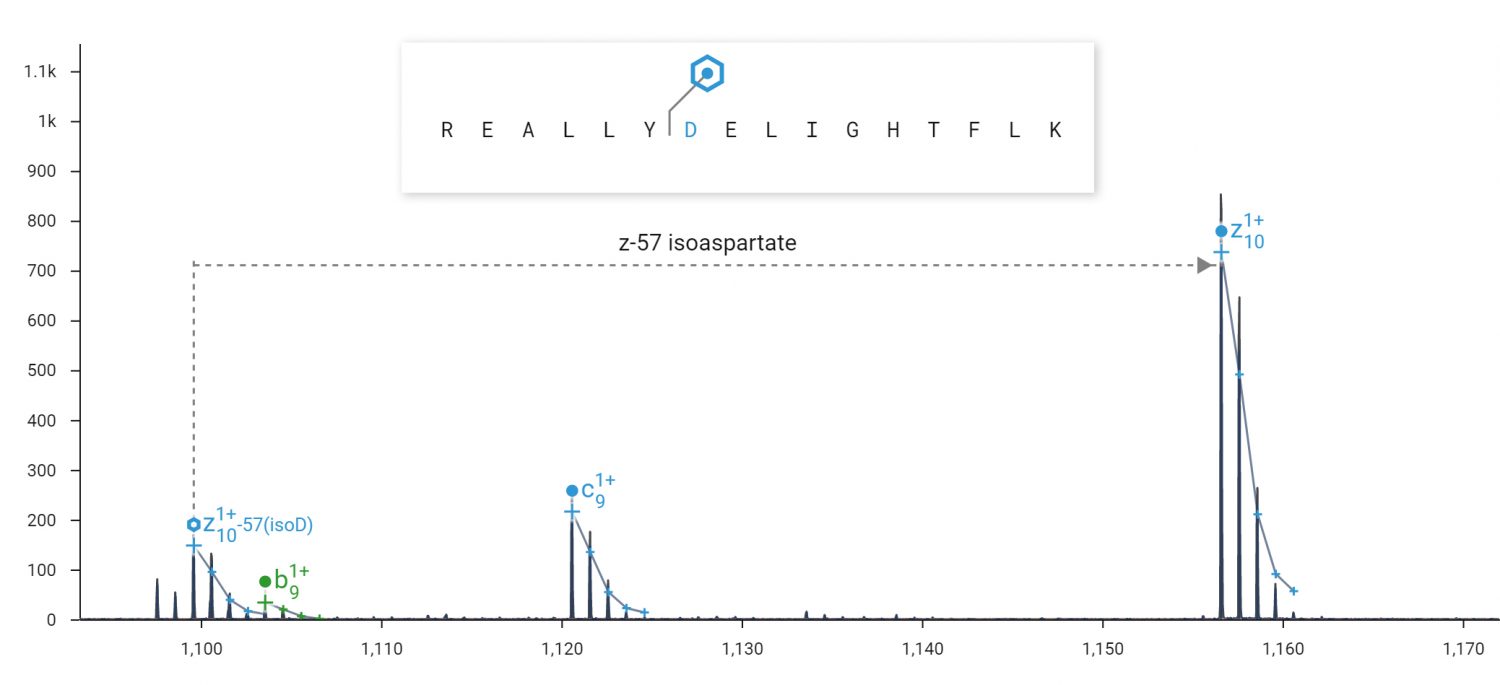
ECD can produce fragment ions diagnostic for isoaspartate. ECD was performed on the 2+ peptide REALLYDELIGHTFLK in an Agilent 6545XT Q-TOF. A 57 Da loss (-C2HO2) from the z10+ ion of the peptide REALLYisoDELIGHTFLK is diagnostic for isoaspartate at position 7. A loss of 45 Da (-CHO2) would be diagnostic for aspartate.
ECD is a “gentle” fragmentation method that preserves labile PTMs – phosphorylation, glycosylation – on fragment ions. With the ExD cell installed, use ECD to map their precise location on peptides and proteins.
Click here to download the Application Note for a demonstration of phosphopeptide analysis using e-MSion’s ExD Cell in an Agilent 6550 iFunnel™ Q-TOF.
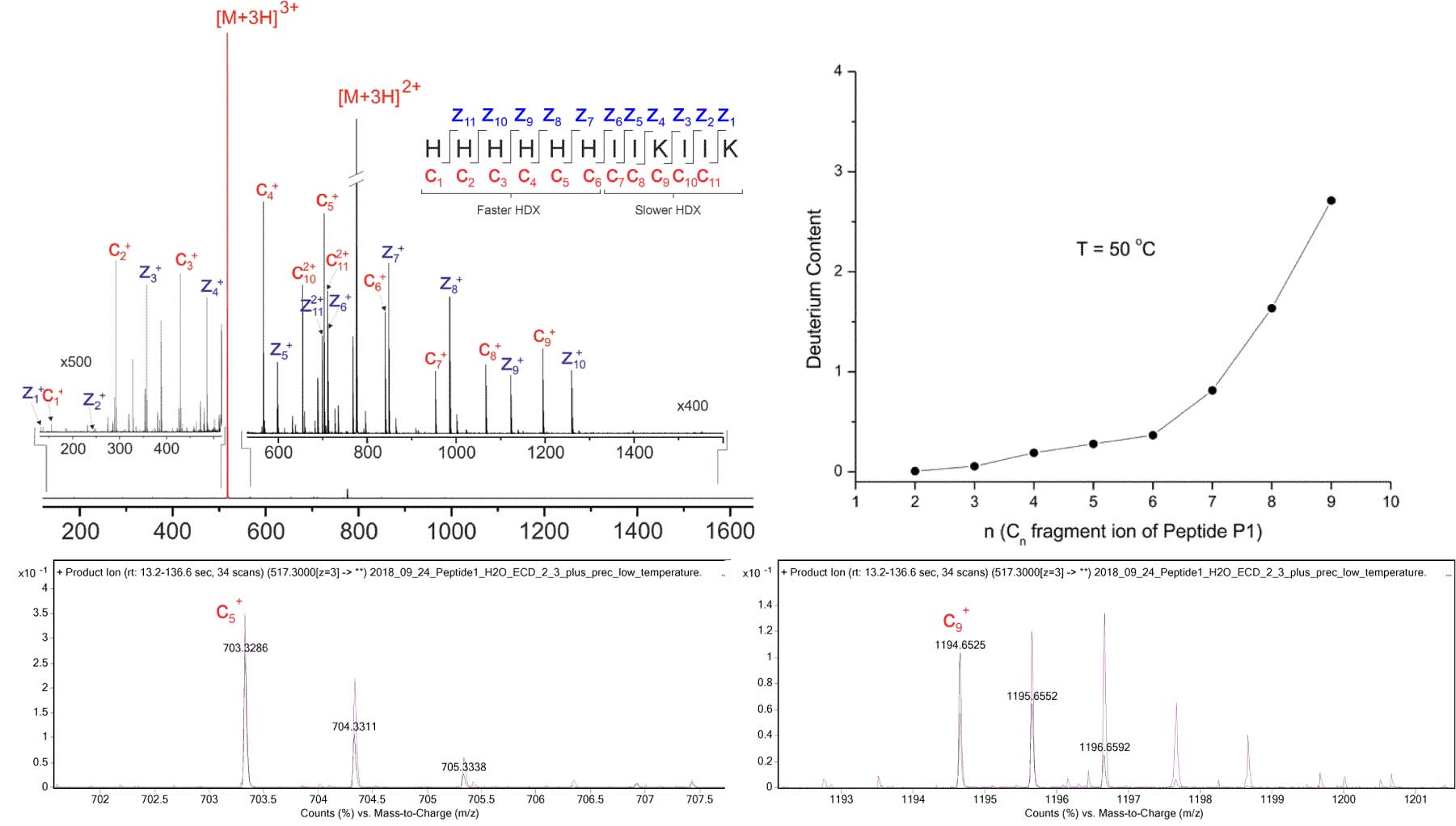
The ExD option enables single-residue-resolution HX-MS with minimal hydrogen scrambling. Top Left: ECD product ion mass spectrum of the [M+3H]3+ precursor of the HX-MS model peptide “P1”, acquired using the ExD Cell in an Agilent 6545 Q-TOF. The His-rich N-terminal half of P1 is engineered to exchange hydrogen more quickly than its C-terminal half. Following back-exchange of deuterated P1, N-terminal c fragments are expected to retain relatively little deuterium, while C-terminal c fragments are expected to retain more deuterium. Any deviations from this pattern would be indicative of hydrogen migration, or “scrambling”, as a result of vibrational excitation. Top Right: After allowing P1 to back-exchange for 5 minutes, deuterium content by residue was similar to expected results published by Rand et al. JACS 130: 1341 (2008): N-terminal residues exchanged hydrogen more quickly than C-terminal residues. Bottom Left: The isotopic distribution of back-exchanged (purple) c5 fragment is similar to protonated (black). Bottom Right: The isotopic distribution of back-exchanged (purple) c9 fragment is shifted right relative to protonated (black), indicating residual deuterium incorporation.
Selective dissociation of disulfide bonds by ECD enables their localization without prior sample reduction.
Figure 1 – Insulin structure and fragments identified in an LC/ECD experiment (average of 5 scans). c– and z-type ECD ions are indicated with blue dots, green dots indicate b– and y-type ions, and purple dots signify a-type ions. Circled dots indicate w-ions. Top: fragments found with Cys7-Cys7 disulfide bridge assumed to be intact. Each chain was calculated separately. The fragments of the B-chain were calculated with the mass of the A-chain as a modification on Cys7, and fragments of the A-chain were calculated with the mass of the B-chain on its Cys7. Bottom: fragments found with the Cys19-Cys20 disulfide bridge intact. The fragments were calculated in the same way as the upper figure, with the mass of the A-chain on Cys19 of the B-chain and the mass of the B-chain on Cys20 of the A-chain.
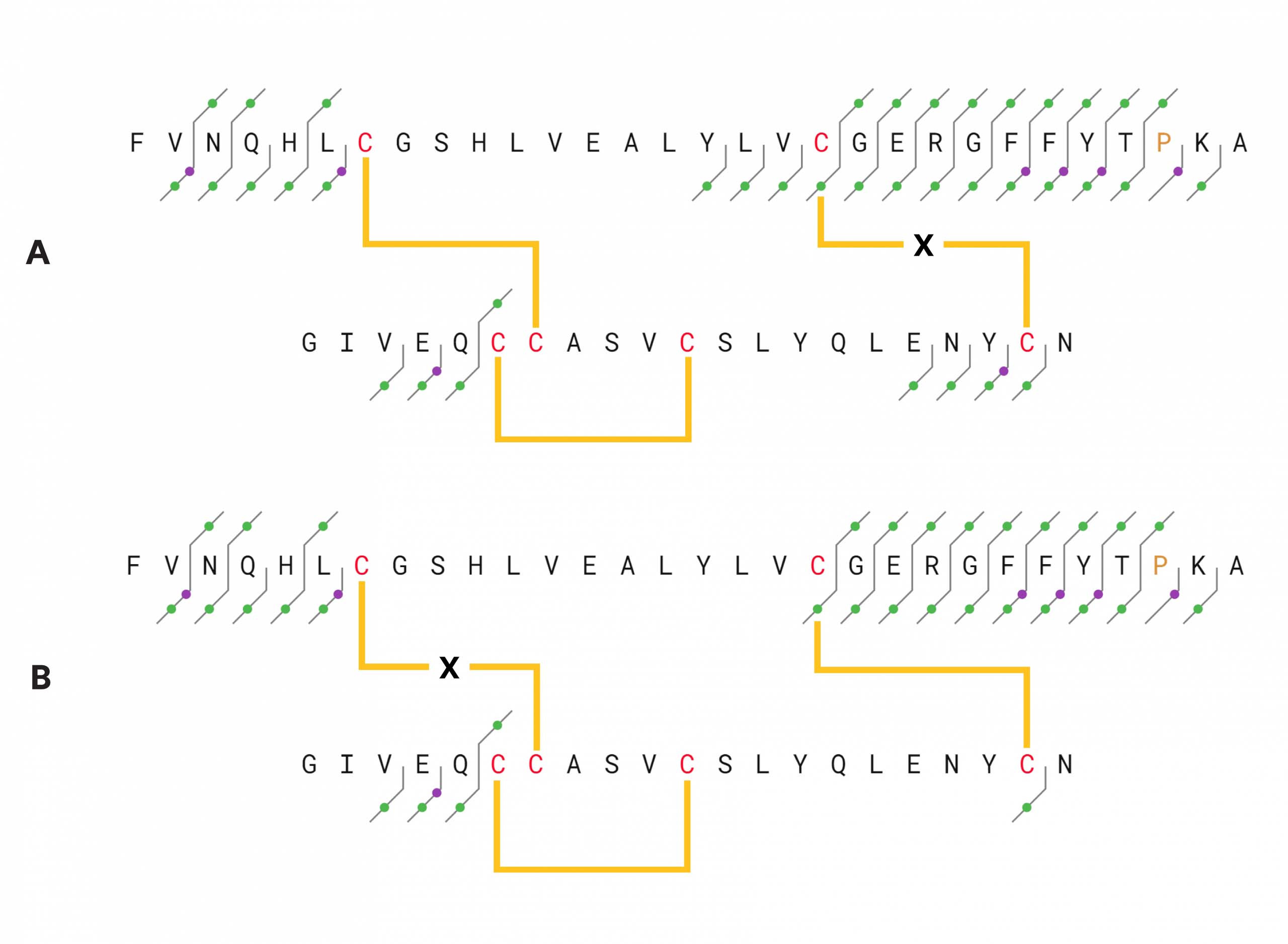 Figure 2 – Insulin structure and fragments identified by CID, 25 V collision energy.
Figure 2 – Insulin structure and fragments identified by CID, 25 V collision energy.
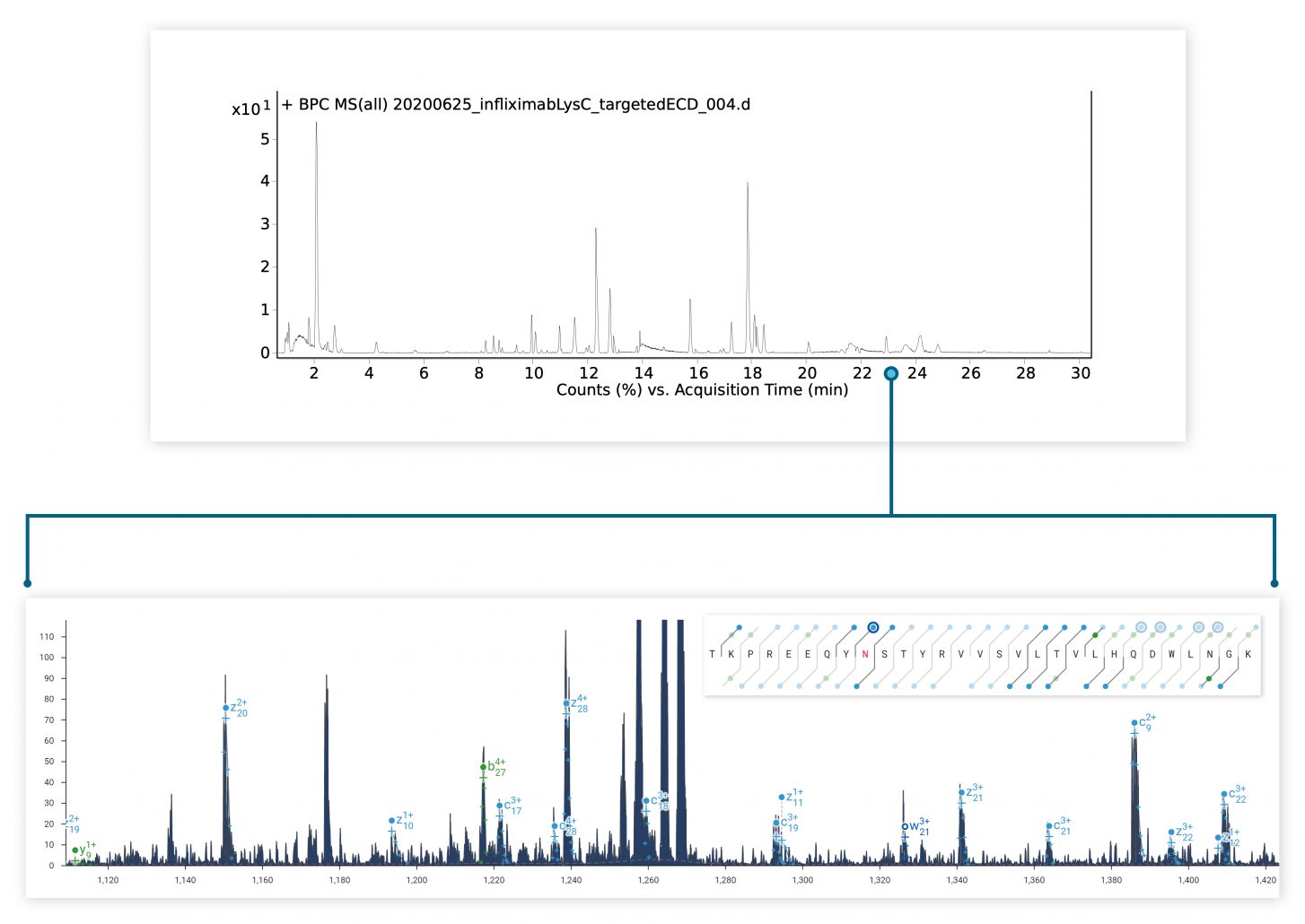
The ExD option is compatible with HPLC timescales and adds valuable information to bottom-up peptide mapping experiments. Here, ECD was used in a targeted manner to fragment a glycopeptide from a Lys C digest of a mAb. The zoomed-in spectrum shows a series of z ions confirming the site & composition of the N glycan without fragmenting the glycan itself.
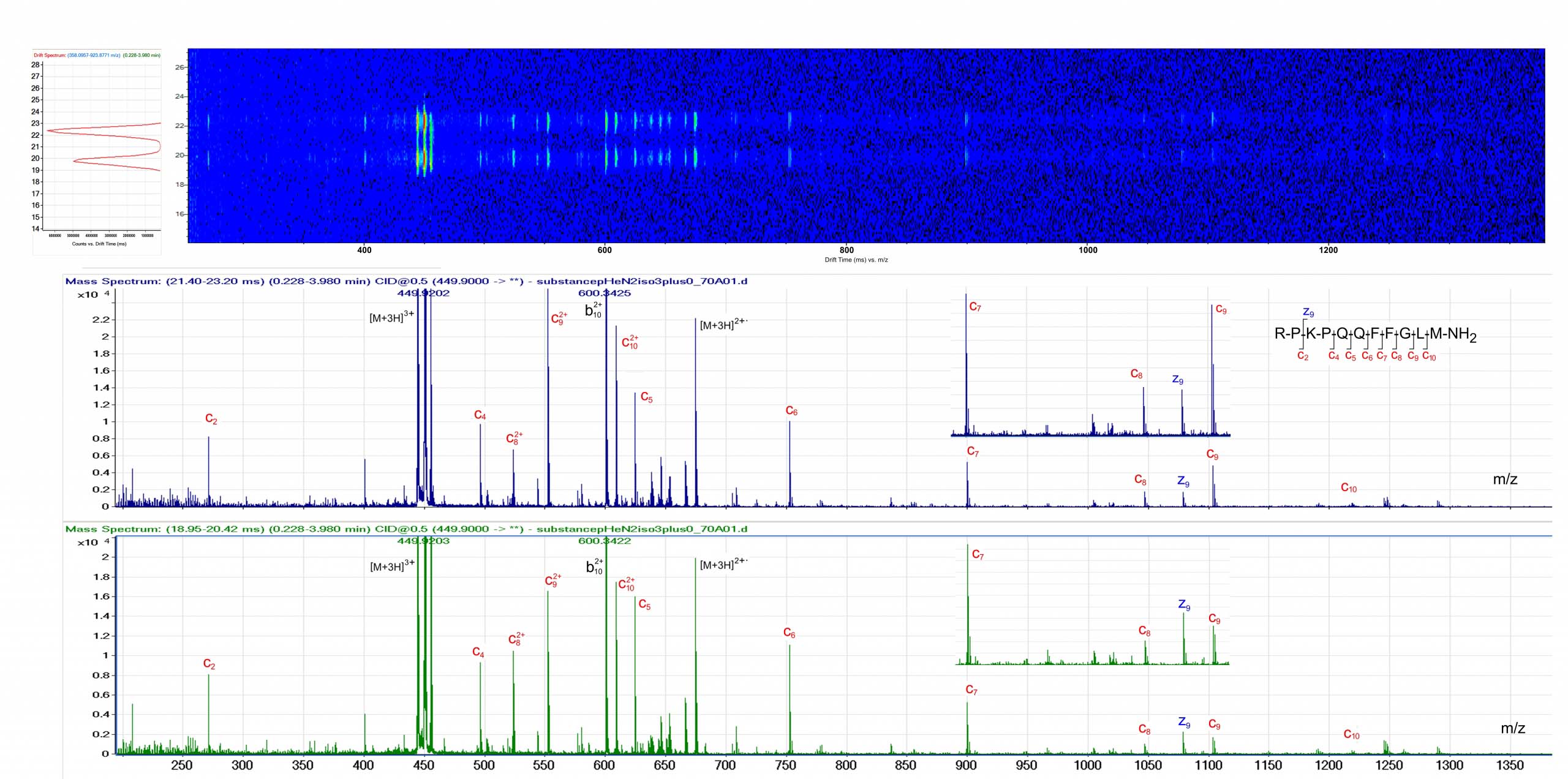 The ExD option is compatible with ion mobility separation in the Agilent 6560 Ion Mobility LC/Q-TOF. Top: Ion Mobility drift spectrum revealing two conformers of [M+3H]3+ Substance P. Bottom: The ExD cell was used to generate ECD product ion mass spectra of the two different Substance [M+3H]3+ conformers. Data courtesy of Dr. Cathy Costello, Boston University.
The ExD option is compatible with ion mobility separation in the Agilent 6560 Ion Mobility LC/Q-TOF. Top: Ion Mobility drift spectrum revealing two conformers of [M+3H]3+ Substance P. Bottom: The ExD cell was used to generate ECD product ion mass spectra of the two different Substance [M+3H]3+ conformers. Data courtesy of Dr. Cathy Costello, Boston University.Discover what you're missing without ExD
Install the ExD cell in your current instrument to maximize your investment, or add ExD to an instrument purchase order to capitalize on the increased sensitivity and resolution of newer technology.
The ExD cell attaches to the entrance of a shortened collision cell, replacing the original collision cell during installation. This way, ExD occurs after isolation by the quadrupole mass filter without affecting collision cell operation.
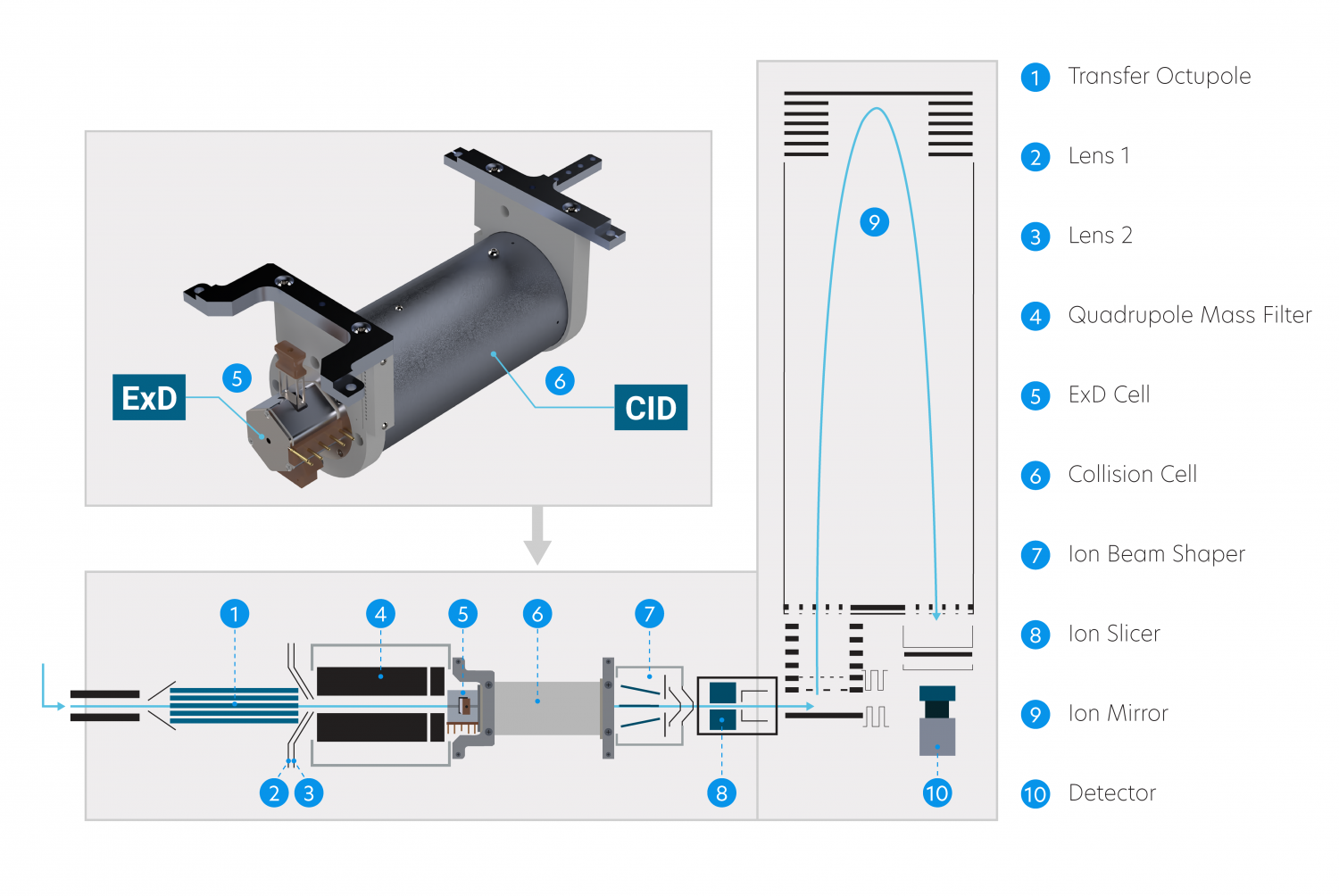
The ExD Option is available for Agilent 6545XT AdvanceBio and 6560 Ion mobility LC/Q-TOF.
Our ExD filament design features a plug-in cassette with exchangeable filament insert for quick and easy replacement.
ECD is a “gentle” fragmentation method that preserves labile PTMs – phosphorylation, glycosylation – on fragment ions. With the ExD cell installed, use ECD to map their precise location on peptides and proteins.
Selective dissociation of disulfide bonds by ECD enables their localization without prior sample reduction.
The ExD cell can operate in either ECD or transmission-only mode.
Now, the ExD Software enables automatic switching between selected ExD modes in sync with changes in instrument scan type during data acquisition.
Use the ExD Software autotune feature to simplify operation of the ExD cell.
The ExD AQ-25x Option for Agilent LC/Q-TOF makes electron-based fragmentation methods accessible and highly affordable for pharmaceutical, clinical, consumer product, and life science protein research laboratories.
Increase sequence coverage of larger peptides and intact proteins beyond the limits of CID alone.
Since ECD and CID produce complementary sequence information, the combination of both methods increases confidence in results.
ECD fragmentation products unique to isoaspartate clearly differentiate it from the non-isomerized form. Isoaspartate is implicated in age-related protein inactivation and aggregation, and reduced efficacy of protein therapeutics.
Side chain fragments (w-ions) produced by ECD can be used to distinguish isobaric amino acids.
User Spotlight
Webinar

Bottom-up and Top-down Disulfide Bond Mapping of Beta-lactoglobulin on a Q-TOF with the Capability to Perform both CID and ECD
Rebecca Glaskin, Ph.D.
Disulfide bonds regulate the structure & function of many proteins, and are often critical for the safety and efficacy of biotherapeutics. It is therefore important to have methods to accurately characterize disulfide bonds. In this talk, Dr. Rebecca Glaskin demonstrates improved disulfide bond mapping abilities by performing ECD and CID fragmentation on a standard protein, beta-lactoglobulin. The experiments were carried out on an Agilent 6545XT LC/Q-TOF equipped with an e-MSion ExD AQ-251 Option. Top-down data analysis was accomplished using ExD Viewer for MS/MS fragment matching and sequence confirmation while bottom-up data sets were analyzed using the Protein Metrics disulfide bond workflow feature.
What's in the box?
Add electron-based fragmentation to your current instrument or include with a new instrument purchase. During field installation of the ExD Option, the combined ExD-collision cell replaces the original instrument collision cell.
Now available for Agilent 6545XT AdvanceBio and 6560 Ion Mobility LC/Q-TOF.
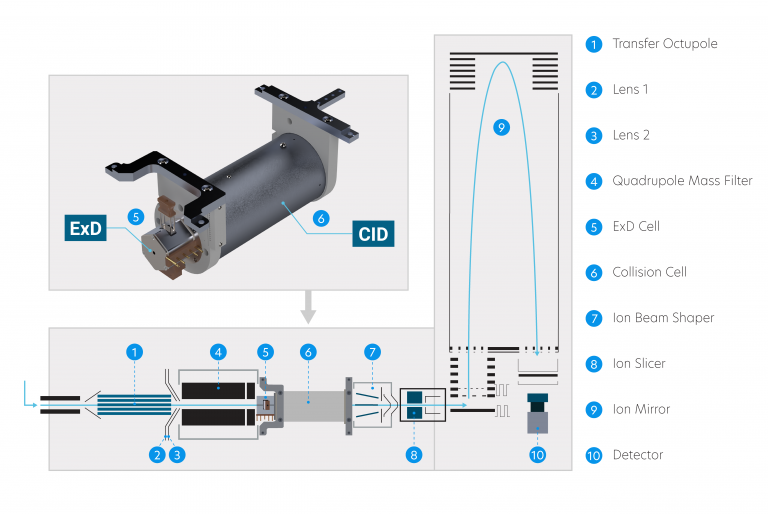


Resources
For Agilent LC/Q-TOF

Download this version of ExDControl if you’re using the ExD AQ-25x Option for Agilent LC/Q-TOF.
FAQ
e-MSion uses “ExD” to refer to the broad range of electron energies the ExD cell can produce. The ExD cell is capable of both low-energy (below 3 eV) electron capture dissociation (ECD) and high-energy (3-20 eV) electron-induced dissociation (EID).
In the literature, “ExD” refers to a family of electron-based fragmentation technologies including ECD, EID, electron transfer dissociation (ETD), electron detachment dissociation (EDD), and negative ion ECD (niECD). EID is also sometimes split into hot ECD (HECD) and electron-impact excitation of ions from organics (EIEIO), depending on the electron energy used.
See Qi and Volmer. 2017. Electron-based Fragmentation Methods in Mass Spectrometry: An Overview. Mass Spectrometry Reviews 36, 4-15.
Including time for unpacking, venting, pump down, and performance verification, a field service engineer will typically require 2 days to complete the installation process. Hardware modification takes about 30 minutes.
Please contact us to learn more about installation requirements.
The ExD cell operates on a microsecond timescale. Its high speed and flow-through design means that the instrument duty cycle is unaffected by installation of the cell.
The ExD cell electron source is a consumable filament, which requires replacement after burning out. Several features in the ExD Software are designed to extend the filament lifespan by protecting it from rapid heat changes.
Filament replacement involves swapping out “plug-in” filament cassettes inside the cell, which minimizes the time spent with the instrument vented and the subsequent pump down time needed before the instrument is operational again. We provide users with instructions for replacing the filament in our product documentation.
Unlike ETD, ECD does not use any reagent. This combined with its distance from the source means the ExD cell does not require regular cleaning.
Use the ExD Software to change ExD cell settings. The same settings for ECD will work on most analytes with minimal adjustment. See Technology for a description of ExD cell operation.
Currently, the ExD AQ-250 Option for Agilent LC/Q-TOF offers an autotune feature for simplified use. Autotuning for our other products is under development.
Due to its flow-through design, the ExD cell operates at a speed compatible with HPLC, UPLC, CE, and IM separation methods.
The ExD cell is designed to fragment polypeptides, but can also be tuned to fragment glycans and lipids.
ECD efficiency theoretically increases with the square of ion charge, making the ExD cell more efficient at fragmenting large, multi-charged proteins. In practice, noncovalent interactions limit the ability of ECD alone to dissociate folded proteins larger than ~30kDa. Denaturing conditions and/or supplemental vibrational excitation can be used to increase ECD efficiency of larger proteins.
On the other end of the spectrum, the ExD cell is capable of fragmenting short peptides, albeit with lower efficiency. For ECD, a minimum precursor charge of 2+ is required to detect product ions because of the charge neutralization that occurs with electron capture.
The ExD cell can also be tuned for EID to target singly charged precursors and non-peptidic samples. This technique is currently only useful for applications where sample quantity is not limited. Development to increase EID efficiency is ongoing.
At some point, you or your service engineer may want to temporarily revert the instrument to its default hardware configuration. The “swap-out” design of the ExD cell makes this possible, since all original parts are preserved during installation.